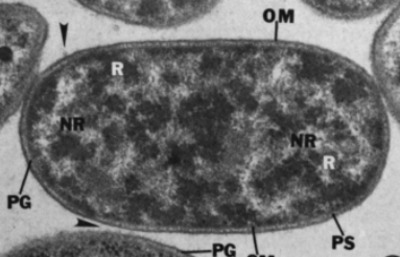
| Species: | Bacteroides fragilis 1 |
|---|---|
| Genus: | Bacteroides |
| Family: | Bacteroidaceae |
| Order: | Bacteroidales |
| Class: | Bacteroidia |
| Phylum: | Bacteroidetes |
| Disease Association: |
Type 1 diabetes (ES=0.4107) Ulcerative colitis (ES=0.378568) |
|---|---|
| Region Enrichment: | Mongolia |
In the linked pathways:
red=enriched, blue=depleted
ko00010 - Glycolysis / Gluconeogenesis
ko00020 - Citrate cycle (TCA cycle)
ko00030 - Pentose phosphate pathway
ko00051 - Fructose and mannose metabolism
ko00061 - Fatty acid biosynthesis
ko00230 - Purine metabolism
ko00240 - Pyrimidine metabolism
ko00250 - Alanine, aspartate and glutamate metabolism
ko00260 - Glycine, serine and threonine metabolism
ko00270 - Cysteine and methionine metabolism
ko00290 - Valine, leucine and isoleucine biosynthesis
ko00300 - Lysine biosynthesis
ko00340 - Histidine metabolism
ko00400 - Phenylalanine, tyrosine and tryptophan biosynthesis
ko00450 - Selenocompound metabolism
ko00471 - D-Glutamine and D-glutamate metabolism
ko00473 - D-Alanine metabolism
ko00500 - Starch and sucrose metabolism
ko00511 - Other glycan degradation
ko00520 - Amino sugar and nucleotide sugar metabolism
ko00521 - Streptomycin biosynthesis
ko00540 - Lipopolysaccharide biosynthesis
ko00550 - Peptidoglycan biosynthesis
ko00620 - Pyruvate metabolism
ko00660 - C5-Branched dibasic acid metabolism
ko00670 - One carbon pool by folate
ko00710 - Carbon fixation in photosynthetic organisms
ko00720 - Carbon fixation pathways in prokaryotes
ko00730 - Thiamine metabolism
ko00740 - Riboflavin metabolism
ko00750 - Vitamin B6 metabolism
ko00770 - Pantothenate and CoA biosynthesis
ko00780 - Biotin metabolism
ko00785 - Lipoic acid metabolism
ko00790 - Folate biosynthesis
ko00970 - Aminoacyl-tRNA biosynthesis
ko00983 - Drug metabolism - other enzymes
ko03010 - Ribosome
ko03030 - DNA replication
ko03060 - Protein export
ko03410 - Base excision repair
ko03430 - Mismatch repair
ko03440 - Homologous recombination
M00001 - Glycolysis (Embden-Meyerhof pathway), glucose => pyruvate
M00002 - Glycolysis, core module involving three-carbon compounds
M00003 - Gluconeogenesis, oxaloacetate => fructose-6P
M00004 - Pentose phosphate pathway (Pentose phosphate cycle)
M00005 - PRPP biosynthesis, ribose 5P => PRPP
M00006 - Pentose phosphate pathway, oxidative phase, glucose 6P => ribulose 5P
M00007 - Pentose phosphate pathway, non-oxidative phase, fructose 6P => ribose 5P
M00008 - Entner-Doudoroff pathway, glucose-6P => glyceraldehyde-3P + pyruvate
M00010 - Citrate cycle, first carbon oxidation, oxaloacetate => 2-oxoglutarate
M00015 - Proline biosynthesis, glutamate => proline
M00016 - Lysine biosynthesis, succinyl-DAP pathway, aspartate => lysine
M00017 - Methionine biosynthesis, apartate => homoserine => methionine
M00019 - Valine/isoleucine biosynthesis, pyruvate => valine / 2-oxobutanoate => isoleucine
M00020 - Serine biosynthesis, glycerate-3P => serine
M00021 - Cysteine biosynthesis, serine => cysteine
M00022 - Shikimate pathway, phosphoenolpyruvate + erythrose-4P => chorismate
M00023 - Tryptophan biosynthesis, chorismate => tryptophan
M00026 - Histidine biosynthesis, PRPP => histidine
M00045 - Histidine degradation, histidine => N-formiminoglutamate => glutamate
M00048 - Inosine monophosphate biosynthesis, PRPP + glutamine => IMP
M00049 - Adenine ribonucleotide biosynthesis, IMP => ADP,ATP
M00050 - Guanine ribonucleotide biosynthesis IMP => GDP,GTP
M00051 - Uridine monophosphate biosynthesis, glutamine (+ PRPP) => UMP
M00060 - Lipopolysaccharide biosynthesis, KDO2-lipid A
M00061 - D-Glucuronate degradation
M00063 - CMP-KDO biosynthesis
M00064 - ADP-L-glycero-D-manno-heptose biosynthesis
M00086 - beta-Oxidation, acyl-CoA synthesis
M00093 - Phosphatidylethanolamine (PE) biosynthesis, PA => PS => PE
M00096 - C5 isoprenoid biosynthesis, non-mevalonate pathway
M00115 - NAD biosynthesis, aspartate => NAD
M00116 - Menaquinone biosynthesis, chorismate => menaquinol
M00119 - Pantothenate biosynthesis, valine/L-aspartate => pantothenate
M00122 - Cobalamin biosynthesis, cobinamide => cobalamin
M00123 - Biotin biosynthesis, pimeloyl-ACP/CoA => biotin
M00124 - Pyridoxal biosynthesis, erythrose-4P => pyridoxal-5P
M00127 - Thiamine biosynthesis, AIR => thiamine-P/thiamine-2P
M00134 - Polyamine biosynthesis, arginine => ornithine => putrescine
M00140 - C1-unit interconversion, prokaryotes
M00144 - NADH
M00149 - Succinate dehydrogenase, prokaryotes
M00153 - Cytochrome bd ubiquinol oxidase
M00157 - F-type ATPase, prokaryotes and chloroplasts
M00159 - V-type ATPase, prokaryotes
M00176 - Assimilatory sulfate reduction, sulfate => H2S
M00308 - Semi-phosphorylative Entner-Doudoroff pathway, gluconate => glycerate-3P
M00346 - Formaldehyde assimilation, serine pathway
M00432 - Leucine biosynthesis, 2-oxoisovalerate => 2-oxoisocaproate
M00525 - Lysine biosynthesis, acetyl-DAP pathway, aspartate => lysine
M00526 - Lysine biosynthesis, DAP dehydrogenase pathway, aspartate => lysine
M00527 - Lysine biosynthesis, DAP aminotransferase pathway, aspartate => lysine
M00535 - Isoleucine biosynthesis, pyruvate => 2-oxobutanoate
M00549 - Nucleotide sugar biosynthesis, glucose => UDP-glucose
M00554 - Nucleotide sugar biosynthesis, galactose => UDP-galactose
M00565 - Trehalose biosynthesis, D-glucose 1P => trehalose
M00570 - Isoleucine biosynthesis, threonine => 2-oxobutanoate => isoleucine
M00573 - Biotin biosynthesis, BioI pathway, long-chain-acyl-ACP => pimeloyl-ACP => biotin
M00577 - Biotin biosynthesis, BioW pathway, pimelate => pimeloyl-CoA => biotin
M00579 - Phosphate acetyltransferase-acetate kinase pathway, acetyl-CoA => acetate
M00627 - beta-Lactam resistance, Bla system
M00632 - Galactose degradation, Leloir pathway, galactose => alpha-D-glucose-1P
M00642 - Multidrug resistance, efflux pump MexJK-OprM
M00704 - Tetracycline resistance, efflux pump Tet38
M00705 - Multidrug resistance, efflux pump MepA
M00718 - Multidrug resistance, efflux pump MexAB-OprM
M00793 - dTDP-L-rhamnose biosynthesis
M00840 - Tetrahydrofolate biosynthesis, mediated by ribA and trpF, GTP => THF
M00843 - L-threo-Tetrahydrobiopterin biosynthesis, GTP => L-threo-BH4
M00844 - Arginine biosynthesis, ornithine => arginine
M00845 - Arginine biosynthesis, glutamate => acetylcitrulline => arginine
M00854 - Glycogen biosynthesis, glucose-1P => glycogen/starch
M00855 - Glycogen degradation, glycogen => glucose-6P
Undetected
Undetected
Other secondary metabolites
MATLAB species model file: msp_0010.mat