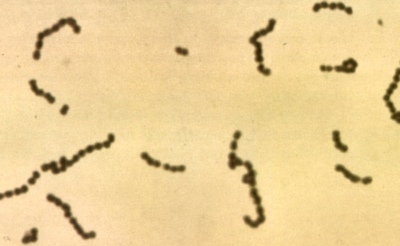
| Species: | Streptococcus salivarius |
|---|---|
| Genus: | Streptococcus |
| Family: | Streptococcaceae |
| Order: | Lactobacillales |
| Class: | Bacilli |
| Phylum: | Firmicutes |
| Gut inflow: | 0.327 |
|---|---|
| Gut outflow: | 0.325 |
| Disease Association: |
Cardiovascular disease (ES=1) NAFLD (ES=1) Type 1 diabetes (ES=0.774052) |
| Region Enrichment: | Westernized (Austria, France, Japan) |
| Shape | Coccus-shaped |
|---|---|
| Gram staining | Gram+ |
| Motility | Nonmotile |
| Oxygen Requirement | Facultative; Anaerobe; Aerobe |
| Sporulation | Nonsporulating |
| Ecosystem | Human; Unclassified; Animal |
| Ecosystem Type | Circulatory system; Skin; Respiratory system; Unclassified; Milk; Excretory system; Digestive system |
In the linked pathways:
red=enriched, blue=depleted
ko00010 - Glycolysis / Gluconeogenesis
ko00061 - Fatty acid biosynthesis
ko00240 - Pyrimidine metabolism
ko00250 - Alanine, aspartate and glutamate metabolism
ko00270 - Cysteine and methionine metabolism
ko00290 - Valine, leucine and isoleucine biosynthesis
ko00300 - Lysine biosynthesis
ko00340 - Histidine metabolism
ko00400 - Phenylalanine, tyrosine and tryptophan biosynthesis
ko00450 - Selenocompound metabolism
ko00471 - D-Glutamine and D-glutamate metabolism
ko00473 - D-Alanine metabolism
ko00500 - Starch and sucrose metabolism
ko00511 - Other glycan degradation
ko00521 - Streptomycin biosynthesis
ko00550 - Peptidoglycan biosynthesis
ko00620 - Pyruvate metabolism
ko00660 - C5-Branched dibasic acid metabolism
ko00670 - One carbon pool by folate
ko00710 - Carbon fixation in photosynthetic organisms
ko00750 - Vitamin B6 metabolism
ko00770 - Pantothenate and CoA biosynthesis
ko00785 - Lipoic acid metabolism
ko00791 - Atrazine degradation
ko00970 - Aminoacyl-tRNA biosynthesis
ko00983 - Drug metabolism - other enzymes
ko03010 - Ribosome
ko03030 - DNA replication
ko03060 - Protein export
ko03410 - Base excision repair
ko03430 - Mismatch repair
ko03440 - Homologous recombination
M00002 - Glycolysis, core module involving three-carbon compounds
M00005 - PRPP biosynthesis, ribose 5P => PRPP
M00007 - Pentose phosphate pathway, non-oxidative phase, fructose 6P => ribose 5P
M00010 - Citrate cycle, first carbon oxidation, oxaloacetate => 2-oxoglutarate
M00015 - Proline biosynthesis, glutamate => proline
M00016 - Lysine biosynthesis, succinyl-DAP pathway, aspartate => lysine
M00017 - Methionine biosynthesis, apartate => homoserine => methionine
M00018 - Threonine biosynthesis, aspartate => homoserine => threonine
M00019 - Valine/isoleucine biosynthesis, pyruvate => valine / 2-oxobutanoate => isoleucine
M00020 - Serine biosynthesis, glycerate-3P => serine
M00021 - Cysteine biosynthesis, serine => cysteine
M00022 - Shikimate pathway, phosphoenolpyruvate + erythrose-4P => chorismate
M00023 - Tryptophan biosynthesis, chorismate => tryptophan
M00026 - Histidine biosynthesis, PRPP => histidine
M00049 - Adenine ribonucleotide biosynthesis, IMP => ADP,ATP
M00050 - Guanine ribonucleotide biosynthesis IMP => GDP,GTP
M00053 - Pyrimidine deoxyribonuleotide biosynthesis, CDP/CTP => dCDP/dCTP,dTDP/dTTP
M00095 - C5 isoprenoid biosynthesis, mevalonate pathway
M00140 - C1-unit interconversion, prokaryotes
M00157 - F-type ATPase, prokaryotes and chloroplasts
M00364 - C10-C20 isoprenoid biosynthesis, bacteria
M00365 - C10-C20 isoprenoid biosynthesis, archaea
M00432 - Leucine biosynthesis, 2-oxoisovalerate => 2-oxoisocaproate
M00525 - Lysine biosynthesis, acetyl-DAP pathway, aspartate => lysine
M00526 - Lysine biosynthesis, DAP dehydrogenase pathway, aspartate => lysine
M00527 - Lysine biosynthesis, DAP aminotransferase pathway, aspartate => lysine
M00535 - Isoleucine biosynthesis, pyruvate => 2-oxobutanoate
M00549 - Nucleotide sugar biosynthesis, glucose => UDP-glucose
M00554 - Nucleotide sugar biosynthesis, galactose => UDP-galactose
M00570 - Isoleucine biosynthesis, threonine => 2-oxobutanoate => isoleucine
M00579 - Phosphate acetyltransferase-acetate kinase pathway, acetyl-CoA => acetate
M00627 - beta-Lactam resistance, Bla system
M00632 - Galactose degradation, Leloir pathway, galactose => alpha-D-glucose-1P
M00705 - Multidrug resistance, efflux pump MepA
M00793 - dTDP-L-rhamnose biosynthesis
M00843 - L-threo-Tetrahydrobiopterin biosynthesis, GTP => L-threo-BH4
M00844 - Arginine biosynthesis, ornithine => arginine
M00845 - Arginine biosynthesis, glutamate => acetylcitrulline => arginine
Undetected
Undetected
Bacteriocin
MATLAB species model file: msp_0380.mat