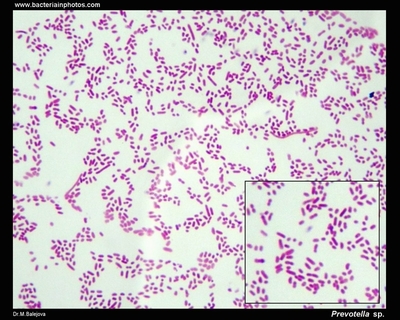
| Species: | Prevotella veroralis |
|---|---|
| Genus: | Prevotella |
| Family: | Prevotellaceae |
| Order: | Bacteroidales |
| Class: | Bacteroidia |
| Phylum: | Bacteroidetes |
| Disease Association: |
Acute diarrhea (ES=0.313173) Liver cirrhosis (ES=1) |
|---|
| Shape | Rod-shaped |
|---|---|
| Gram staining | Gram- |
| Motility | Nonmotile |
| Oxygen Requirement | Anaerobe |
| Sporulation | Nonsporulating |
| Ecosystem | Human |
In the linked pathways:
red=enriched, blue=depleted
ko00020 - Citrate cycle (TCA cycle)
ko00061 - Fatty acid biosynthesis
ko00240 - Pyrimidine metabolism
ko00250 - Alanine, aspartate and glutamate metabolism
ko00300 - Lysine biosynthesis
ko00471 - D-Glutamine and D-glutamate metabolism
ko00473 - D-Alanine metabolism
ko00511 - Other glycan degradation
ko00521 - Streptomycin biosynthesis
ko00540 - Lipopolysaccharide biosynthesis
ko00550 - Peptidoglycan biosynthesis
ko00670 - One carbon pool by folate
ko00710 - Carbon fixation in photosynthetic organisms
ko00730 - Thiamine metabolism
ko00750 - Vitamin B6 metabolism
ko00770 - Pantothenate and CoA biosynthesis
ko00780 - Biotin metabolism
ko00785 - Lipoic acid metabolism
ko00970 - Aminoacyl-tRNA biosynthesis
ko00983 - Drug metabolism - other enzymes
ko03030 - DNA replication
ko03060 - Protein export
ko03430 - Mismatch repair
ko03440 - Homologous recombination
M00005 - PRPP biosynthesis, ribose 5P => PRPP
M00020 - Serine biosynthesis, glycerate-3P => serine
M00021 - Cysteine biosynthesis, serine => cysteine
M00050 - Guanine ribonucleotide biosynthesis IMP => GDP,GTP
M00051 - Uridine monophosphate biosynthesis, glutamine (+ PRPP) => UMP
M00060 - Lipopolysaccharide biosynthesis, KDO2-lipid A
M00063 - CMP-KDO biosynthesis
M00086 - beta-Oxidation, acyl-CoA synthesis
M00093 - Phosphatidylethanolamine (PE) biosynthesis, PA => PS => PE
M00096 - C5 isoprenoid biosynthesis, non-mevalonate pathway
M00116 - Menaquinone biosynthesis, chorismate => menaquinol
M00119 - Pantothenate biosynthesis, valine/L-aspartate => pantothenate
M00140 - C1-unit interconversion, prokaryotes
M00144 - NADH
M00149 - Succinate dehydrogenase, prokaryotes
M00153 - Cytochrome bd ubiquinol oxidase
M00157 - F-type ATPase, prokaryotes and chloroplasts
M00525 - Lysine biosynthesis, acetyl-DAP pathway, aspartate => lysine
M00526 - Lysine biosynthesis, DAP dehydrogenase pathway, aspartate => lysine
M00527 - Lysine biosynthesis, DAP aminotransferase pathway, aspartate => lysine
M00554 - Nucleotide sugar biosynthesis, galactose => UDP-galactose
M00565 - Trehalose biosynthesis, D-glucose 1P => trehalose
M00579 - Phosphate acetyltransferase-acetate kinase pathway, acetyl-CoA => acetate
M00632 - Galactose degradation, Leloir pathway, galactose => alpha-D-glucose-1P
M00704 - Tetracycline resistance, efflux pump Tet38
M00705 - Multidrug resistance, efflux pump MepA
M00718 - Multidrug resistance, efflux pump MexAB-OprM
M00793 - dTDP-L-rhamnose biosynthesis
M00855 - Glycogen degradation, glycogen => glucose-6P
Undetected
Undetected
Undetected
Undetected
MATLAB species model file: msp_1808.mat